 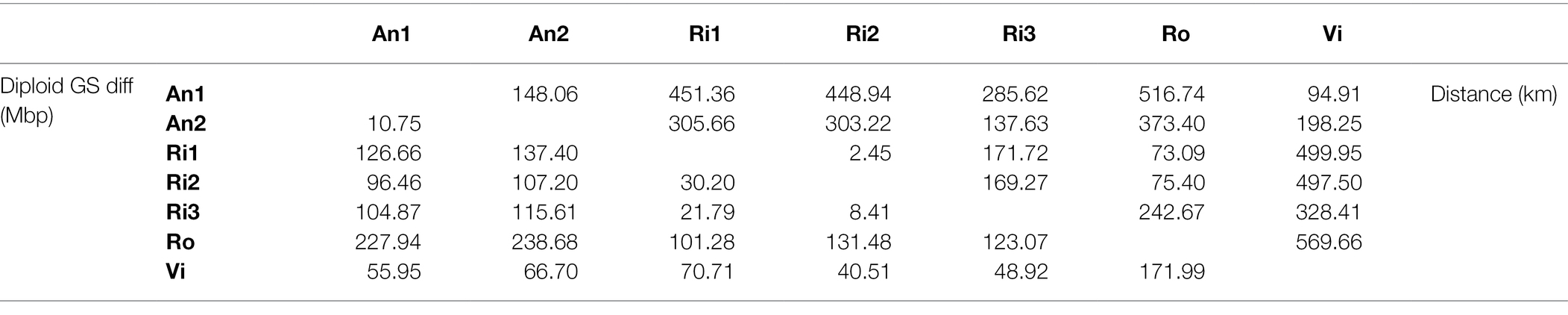
Geographical distances were estimated as the shortest distance by sea between the locality pairs as determined by Google Earth. gliroides were significantly correlated with geographic distances. Euclidean spatial distances and matrix comparisons with 10,000 permutations were performed using NTSYS-pc v2.0. These results suggest that some of the D. The six microsatellite loci yielded a total of 146 alleles, ranging from 1442 per locus. (You can view the original function with: findMethods(ecogen2gstudio).Gliroides populations would have survived in glacial refuges, with posterior expansions after ice retreat. The following chunk contains code adapted from within the ecogen2gstudio function, tweaked to work for our data. This should be easy with the function EcoGenetics::ecogen2gstudio. We will also use some functions from the package gstudio, hence we import the individual-level genetic data into gstudio. The object ‘Frogs.genpop’ has 30 rows, each representing a population. However, this can create warnings later on when calculating genetic distances. Note: Alternatively, we could directly import the ecopop object into a genpop object ( adegenet) with EcoGenetics::ecopop2genpop(Frogs.ecopop). # adegenet::genind2genpop(x = Frogs.genind) # locus factor for the 43 columns of list of allele names for each locus # number of alleles per locus (range: 3-9) # // 30 populations 8 loci 43 alleles size: 18.2 Kb In addition, EcoGenetics created a table of absolute frequencies (i.e. counts) of alleles for each individual in slot A.įrogs.genpop # /// GENPOP OBJECT ///////// The summary confirms that we now have data in the S and G slots. # | slot S: | -> 413 x 2 structures > 2 structures found # | slot E: | -> 0 x 0 environmental variables # | slot G: | -> 413 x 8 loci > ploidy: 2 || codominant # | slot P: | -> 0 x 0 phenotypic variables Linking paternity to ecological variables Nested model (NMLPE) for hierarchical sampling designs A quick note on controlling for population structure in RDA Redundancy Analysis (RDA): a multivariate GEA Latent Factor Mixed Models (LFMM): a univariate GEA How does changing resolution affect these metrics? Convert conductance into effective distance Setting cost values and calculating conductance Simulated data: 2-island model with admixture Benchmarking file import and export options Run simulator using a previously defined parameter set Calculate Hanski’s index Si with source patch parameters Fit spatially varying coefficients model with package ‘spmoran’ Spatial filtering with MEM using package ‘spmoran’ 
Fit spatial simultaneous autoregressive error models (SAR) Fit models with spatially correlated error (GLS) with package ‘nlme’ Test regression residuals for spatial autocorrelation 
Assess correlation between trait and environment Estimate genetic and non-genetic variance components from a common garden experiment Specify spatial weights and calculate Moran’s I Create Mantel correlogram for genetic data Are genetic differentiation and diversity related? What determines genetic differentiation among sites? Spatial distribution of genetic structure Aggregate genetic data at population level (allele frequencies) Basic checking of markers and populations Use ‘terra’ and ‘tmap’ to display categorical map with color scheme Convert ‘SpatialPointsDataFrame’ to ‘sf’ object Sample landscape metrics within buffer around sampling locations Display raster data and overlay sampling locations, extract data View information stored in ‘genind’ object 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
